Neuropype Experiment Recorder (beta)
Overview
Neuropype Experiment Recorder (ER) is a cognitive task presentation (similar to PsychoPy) and signal data recording application combined into one.
- It is designed to be simple and easy to use.
- It runs task modules written in Python using an extension of the open-source Simulation and Neuroscience Application Platform (SNAP) framework. ER ships with a number of standard cognitive tasks, and you can fairly easily write your own based on the sample tasks provided, with minimal coding depending on the complexity of the task.
- It can be used to record biosignal data from any device(s) that support LSL, including EEG, fNIRS, ECG, EMG, GSR, EyeTracking, Accelerometer, etc.
- For signal data capture and recording, it uses Neuropype to capture one or multiple Lab Streaming Layer (LSL) streams (or directly connect to certain supported EEG devices), and saves these to the open-source XDF file format (which allows multiple signal and event streams to be saved in the same file), along with timesynced event markers produced by the task(s) themselves.
- It is highly configurable.
- Individual cognitive tasks can be configured using simple text configuration files, and you can have multiple different configuration files which reference the same task with different presets (i.e., different groups, or for testing, etc.).
- It can run a sequence of cognitive tasks (i.e, a multi-task study) by simply listing the tasks to run in a study settings file.
- Settings for a given study (i.e., tasks to run, LSL streams to capture, etc.) can be saved in individual study settings files which can be launched as needed.
A recording session can be configured to run several different tasks in succession, with each task being saved to an appropriately named XDF file. A study settings file (with an .er extension) can specify the task configurations to run as well as the LSL stream(s) to capture, and other presets.
ER is ideal for simultaneously capturing data from multiple devices (i.e., EEG, cardiac, physio, GSR, eye tracking, etc.) and saving these to a single data file along with event markers, for multi-modal analysis in Neuropype (or other apps).
Since Experiment Recorder uses Neuropype under the hood, it comes with a real-time signal quality pipeline for checking signal quality before starting a session. Optionally, you can use ER to run a Neuropype pipeline for real-time mapping of cortex activity and connectivity flow during a task or play back a recorded session. These pipelines can be used to monitor signal quality or view brain activity on a second computer running ER and configured to capture the same LSL streams as the computer where the tasks are being conducted.
For Neuroscale users, ER can automatically upload your session's XDF files to your Neuroscale storage.
Experiment Recorder is in beta, and provided as part of the Neuropype Suite under the terms of the Neuropype license.
Installation
- ER is installed along with your Neuropype Suite, and will be in an
Experiment Recorderfolder in your installation (i.e.,c:\Program Files\Neuropype Academic Suite\Experiment Recorder). - An
ERdesktop shortcut will be created on install. - The
.erfile extension will be associated with Experiment Recorder, allowing you to launch ER with a set of presets, by simply double-clicking on your.erstudy file. (See below.) - If during installation or first running ER, a Windows Defender window pops up asking you to allow Python to communicate on the network, you should grant such permission. This is necessary for LSL connections to work over the network and for the ER app to communicate with Neuropype.
Launching Experiment Recorder
- First, Neuropype itself must be running. Neuropype runs as a background service, and
you will see its
NPicon in your Windows system tray if it is running. (Neuropype will automatically launch on Windows startup.) It uses very little resources when idle so you can keep it running all the time. If you ever do need to quit Neuropype, right-click on theNPicon and selectExit Neuropype. - Launch ER either through the Experiment Recorder desktop shortcut, or by double-clicking an
.erstudy file pre-configured for your study. (See Study Design Files below.) - The ER interface is intentionally very basic in order to simplify the task of the
experiment operator and avoid errors. All study configuration is done in a plain text study (
.er) file, and all configuration of individual tasks are done in their corresponding plain text task config (.cfg) files. If you need different configurations for different groups or individuals in a study, you can create separate.erfiles which in turn refer to specific.cfgfiles with the corresponding settings for those tasks for a given
individual or group. - The study files (
.er) and task config files (.cfg) are plain text files that can be opened and edited with any text editor, such as the Notepad app that comes with Windows. (Do not edit them in Microsoft Word.)
Command line launcher
- ER can also be launched from the command line with the
experiment-recorder.cmdWindows command file located in yourExperiment Recorderinstallation folder. This allows you to invoke ER from another script or application (you can also invoke the python command in theexperiment-recorder.cmdfile, just be sure that you're invoking it using the python interpreter that ships with ER and is located in theExperiment Recorder/pythonfolder.)
Running a session
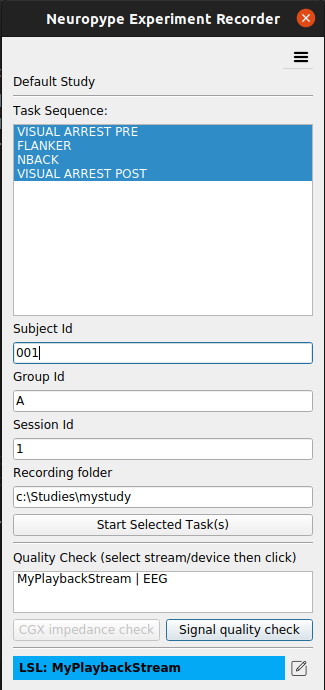
- When you Launch Experiment Recorder, the following window will appear:
- A list of tasks will be shown, which are the tasks specified in the [Tasks] section of
the
.erstudy file that was run. They will be executed in the order shown, one after the other. (To change this order, modify the order of the lines in the.erfile.) (More on tasks below under Study Settings Files). - Enter the subject id, and an optional session id.
- The bottom section of the ER Window will show the LSL Stream(s) (or EEG headset) that ER is presently configured to listen for. You can click the icon to the right of the blue Status Bar to select a different LSL stream or device. See LSL Streams below for details.
- Optionally run the Signal Quality Check by clicking the corresponding button (if using a CGX headset you can do also perform an Impedance Check by clicking that button.)
- By default, all the tasks listed in the
.erfile are selected. You can optionally deselect tasks that you don't want to run for this session. This allows you to, for example, repeat only one task in the set for a user due to a malfunction or interruption. - Click Start Task to begin the selected tasks.
- The ER window will close and a separate task window will appear where text from the task will be displayed and subject input (i.e., keyboard input) will be captured.
- The bottom bar of the ER window shows a status/activity bar. On startup, this shows the expected source and LSL stream (if the source in your
.erfile is 'lsl'), or the device name (if the source in your.erfile is 'cgx'). If you notice that this doesn't match your expected setup, modify the [Input] section of your .er file, or click the icon next to the status bar to select a different LSL stream or device. Your LSL stream or device name must be correct to record data. (See also LSL Streams below.) - When the tasks complete and the files have been uploaded to Neuroscale (if that option is enabled), the main window will reappear. To exit ER, simply close the window (by clicking the usual close window icon), or, to run the same session again with a different subject, simply change the SubjectID, GroupID and SessionID as usual, make sure the correct tasks are selected, and click Start Selected Tasks. (However, in most cases you'll want to first run the Signal Quality Check with the headset on the new subject.)
- You can cancel a session at any time by pressing the Escape key in the task window. This will abort the task and terminate ER (Neuropype will continue to run in the background). Any data that has been recorded so far is not lost, as data is saved to disk in realtime. To start over, double-click your study
.erfile to re-run ER.
Default settings
- If launch Experiment Recorder using the desktop shortcut, or you don't specify any tasks in your .er study file
(the [Tasks] section is empty), a dropdown will appear with the list of all the task config
files found in the
configsfolder of your ER installation (i.e.c:\Program Files\Neuropype Academic Suite\Experiment Recorder\configs). In that case, select the desired task to run from the list, and click Start Tasks. In this case, a single task will be run. To run several tasks in sequence, list them in your.erfile and run ER by clicking that file. - The default settings that ER runs with when launched without specifying an
.erfile are in adefault.erfile located in thesettingsfolder of your Experiment Recorder installation. You may also edit this file to change the default behavior, such as the recording folder. You can also use this file as a template for creating your own study-specific study files (see below).
Study settings files
- Each study has study settings file with an
.erfile extension, that defines the Experiment Recorder settings for the study. .erfiles are plain text files that you can open and edit with any text editor such as Notepad.- ER ships with a default
.erfile located in yourExperiment Recorder/settingsfolder of your installation. You can copy and use this as a template for your own study files. It includes comments throughout (using#as the first character in the line) to describe the contents. - By default, ER will look for study files in the
Experiment Recorder/settingsfolder of your installation. However, an.erfile can be located and launched from anywhere on the same drive. - There three ways to launch ER with a specific set of settings:
- Double-click the
.erfile (unless you removed the file association or another program you installed overwrote that association). -
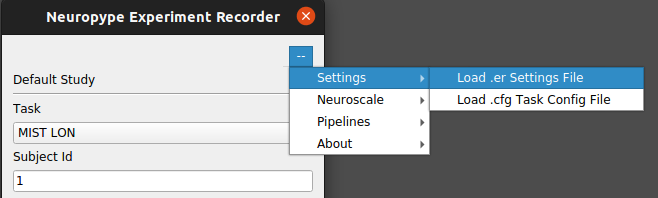
Use the
Settings | Load .er Settings filemenu item and select the desired.erfile to load.
-
Invoke ER from the command line using the
experiment-recorder.cmdWindows command file and pass the .er file with the--settingsargument. I.e.,experiment-recorder.cmd --settings=c:\MyStudies\study2.er(This is particularly useful for launcher ER from another script or app.) - Study settings files (
.er) use theINIconfiguration file format (see ). This is a simple format that is easy to read and write. Thesettings/default.erfile has comments throughout that explain the various settings. It is very important that you do not modify the structure of the file or the section headings or the settings will not load properly. -
Study files contain the following sections:
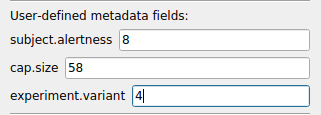
[Input]: Defines whether the source is LSL or CGX, and which LSL streams are to be captured. (Presently only CGX (Cognionics) headsets are supported for direct connect with the need to create an LSL stream; for other headsets, create an LSL stream using the manufacturer's application or an application available at www.labsteaminglayer.org.)[Output]: Settings that affect the data output to disk, including the path where to save the data, as well as certain settings which are used to determine the filenames (namely, subject_id, group_id, and session_id).[Study]: Several general settings related to the session, including adry_runsetting that allows you to run a cognitive task without recording any data (i.e., for testing).[Metadata]: In addition to the standardsubject_id,group_idandsession_iddefined by default, you may define any other custom metadata tags, and these will be stored in the XDF file along with the signal data. (i.e.hair.color,cap.size, etc.) The tags defined in this section will appear in the ER window as text fields, which the operator can fill in before running a given session. Default values shown in the ER window may also be defined here. See the comments indefault.erfor more information. These values are stored under themetadataproperty of the signal stream in the XDF file. (If you plan to keep your signal data anonymized, do not store PII here.)
[Window]: Settings that define the window where the cognitive task is shown (not the ER dialog). These allow you to change the placement and size of the task window. During installation ER will be configured to display the task window the size of your full screen resolution (i.e., 1920x1280). In some cases you may want to change that to show the window on a second screen (change thewindow_positionsetting to a positive or negative value depending on how your screens are arranged), or to make it smaller so that it is not fullscreen (change thewindow_sizesetting.)[Neuroscale]: If you have a Neuroscale account, you may specify your private storage credentials here and your XDF files will be automatically uploaded to your Neuroscale storage ifupload_to_neuroscaleis set totrue.[Tasks]: These are the tasks to run, defined as<display_name>: <config_file_name>where<display_name>is the name of the task to display in the ER window (and which will be used to name the XDF file for that task), and<config_filename>is the name of a.cfgfile (see Task Config Files below) that defines a cognitive task. I.e.,Pre Visual Arrest: visual_arrest. It is not necessary to specify the.cfgextension. By default, ER will look for the config file in theExperiment Recorder/configsfolder of your installation. However, you may also specify a full path to a config file located elsewhere (i.e.,Pre Visual Arrest: C:\MyStudies\Study1\Tasks\my_va_task.cfg)).- Each setting must be in the format
setting: value. Strings do not need to be enclosed in quotes. This file uses the same syntax as standard .INI files. - Any line that starts with # is a comment line and will be ignored. You may add additional comment lines with notes for your own reference as you wish.
- Do not remove any settings or section headings. You may comment out any settings which are unused by adding a
#to the beginning of that line. - Empty lines are allowed and ignored.
- It can be useful to make a different .er file for each variant of a study, which in turn references different .cfg files with different variations of a task (i.e., with different params or in a different language).
- .er files can be stored anywhere and are launched by double-clicking on them (the
.erfile extension is associated with the ER windows desktop shortcut on installation). They can also be launched from the command line usingexperiment-recorder.cmd --settings=c:\path\to\file.er(experiment-recorder.cmd c:\path\to\file.erwill also work. )
Task config files
- Tasks are written in python and stored in
Experiment Recorder\src\modules(i.e.,flanker.pyc). - Each task to be executed will have a configuration file with a
cfgextension. - Task config files contain settings for a given task that are specific to your study. You may have as many different task config files for a single task as desired.
- By default, task config files are located in the
configsfolder of your ER installation, but can also be kept elsewhere and referenced by their full path in your study.erfile. - A Task Config file is a plain text file that can be edited with any text editor such as Notepad.
- To run a task in ER, you load the task config file, which in turn references the task
module(containing the python code for the task). - The very first line of the config file MUST be the name of the task module. Nothing else should be on that line.
- The remaining lines contain task parameters (settings) which can be modified as desired. For example, the number of trials, trial duration (ITI), volume level, key configurations, sound files, etc. The parameters themselves are defined in the task module.
- If you modify a task's parameters in a
.cfgfile, restart ER for the changes to take effect. You do NOT need to restart Neuropype. - Never change the name of a parameter, only its value, or else the task will fail.
- Each line can contain a single parameter only.
- Task parameters can be commented out (by placing a
#at the start of the line) or deleted altogether. In that case, the module's default value for that parameter will be used. (It is recommended to comment out rather than delete parameters so that you have a record of the parameter name.) - Any line that starts with
#is a comment line and will be ignored. You may add comment lines with notes for your own reference as you wish. - Empty lines are allowed and ignored.
- You can have multiple versions of a task config file for different languages, and then your .er study file can refer to the desired variant/language.
Task parameters
- Task config files use Python syntax to describe task parameters, where each line has:
parameter_name = parameter_value. - Follow the same syntax for each parameter already shown (i.e., if the value is in quote marks, enclose the replacement value in quote marks.)
- String values must be in 'quote marks', numeric values should not.
- If a parameter is a list of values, it must be enclosed in square brackets [].
- String values can span multiple lines by enclosing them in triple double-quotes
"""like this""". This is mostly useful for onscreen text where you want it to be displayed on several lines. The line breaks will be respected (so you can control which words go on each line). Any indentation at the beginning of each line will be ignored (it's just used to make it more readable in the config file). - Parameters can refer to other parameters. For example, you might have
something like this:
response_time = stimulus_duration + interstimulus_intervalIn such case,stimulus_durationandinterstimulus_intervalmust be parameters defined earlier in the same file. - Parameter names are case sensitive.
- Some params may refer to a key to be pressed by the user for a certain action. When specifying a parameter for accepted keystrokes in a task, use the following naming rules:
- Keys that type a character are named that character. It is always lowercase, even when shift or caps lock is pressed. I.e., "a", "b", "3", "[", etc. not "A", "B", "#", "{"
- Keys that don’t type a character are labeled as follows: "escape", "f"+"1-12" (e.g. "f1","f2",..."f12"), "print_screen", "scroll_lock", "backspace", "insert", "home", "page_up", "num_lock", "tab", "delete", "end", "page_down", "caps_lock", "enter", "arrow_left", "arrow_up", "arrow_down", "arrow_right", "shift", "lshift", "rshift", "control", "alt", "lcontrol", "lalt", "space", "ralt", "rcontrol"
- If the text to be shown is in a non-Latin language, be sure to set
the font parameter in the .cfg to the desired non-Latin font. You can add your own
fonts to the
/src/media/fontsfolder of your installation, and they will be found automatically (just specify the font file name, not the whole path).
Task modules
The task modules consist of Python code and these define the structure and flow of a task.
These are located in the /src/modules folder. Each task config file must reference the
task module (.py or .pyc file) to be run, on the very first line of the config file.
Modules use the open-source Simulation and Neuroscience Application Platform (SNAP) framework. SNAP itself uses Panda3D, an open-source game engine, for display functions.
Writing a module does require an understanding of Python, but a simple module isn't too complicated to write.
The source code for a number of task modules are included in the /examples folder of your installation, and
these are fairly well commented, so can be used as a guide for adapting existing tasks shipped with Neuropype or
writing your own.
If you have questions on writing tasks, please pose your questions in the Neuropype Discussion Forums.
Signal Data Input Sources
Lab Streaming Layer (LSL) Streams
- Experiment Recorder records signal data sent over the Lab Streaming Layer (LSL) protocol. See labstreaminglayer.org for more information about LSL.
- The LSL streams to be captured during the task are configured in the
[Input]section of the.erfile. - You can configure up to four different LSL streams in your
.erfile (namedlsl_stream,lsl_stream_two,lsl_stream_three.lsl_stream_four). - LSL is by default network aware. This means you can have your EEG/other device connected to a different
computer than the one ER is running on, generate an LSL stream on that computer, and ER will still be able
to pick it up, and merge it with the event markers being generated by ER itself. However, if you happen
to be running multiple instances of ER on the same LAN at the same time, this could result in one
instance of ER discovering and using the stream being generated on another computer.
In such a case, you can restrict LSL stream discover to the computer on which ER is running by
setting the
lsl_localhost_onlyproperty in the[Input]section of your.erfile. - If necessary you can specify the type of the LSL stream in the
.erfile, which is used to match the LSL stream. This is optional and not usually necessary. - Besides specifying the LSL streams in your
.erfile, you can also select the LSL streams to capture from the ER window before starting a session. Click the icon in the lower-right hand corner next to the blue status bar. A window will appear showing the active LSL streams found on the computer and/or your LAN. You can select one or more LSL streams to record, click Set Selected, and ER will be configured to listen for and record those streams during the session. Note that this is for the current session only. The next time you run ER the settings in your.erfile, or the default settings, will be used.
EEG headsets from CGX
- If you use an EEG headset from CGX (Cognionics) you can specify the model in your
.erfile, or select it manually in the same way as described above for the LSL streams (by clicking the icon next to the blue bar). ER will connect directly to the CGX headset without the need to run the CGX data acquisition app and create an LSL stream.
Session data
- Data is saved to the open-source XDF format, a multi-modal format that can
store multiple data streams (i.e., from different devices) of different modalities (i.e., EEG + eye tracking)
in the same data file. The events from the task modules are also stored in the same XDF file in a
markersstream. - Each task is saved in a separate XDF file. So if your session comprises 3 tasks, 3 separate files will be created for that session.
- Each task file is automatically as follows:
subject-id_task-name_date-time_session_id.xdf, wheresubject-idis the SubjectID specified in the ER window,task-nameis the name of the task as it appears in the ER window (specified in the.erfile under[Tasks]),date-timeis a string representation of the start time of the task (DD-MM-YY:HH-MM-SS) andsession-idis the SessionID specified in the ER window.
Session logs
- The logs for each ER session (both log messages from ER itself and those generate by each task) are saved in a .log file named with date and time of when the session begun (i.e.,
session-2022-11-21_10-32-13.log). Log files are stored in your user folder under.er/logs(i.e.,C:\Users\myuser\.er\logs) - Log messages can be seen in realtime by invoking ER using
experiment-recorder.cmd /console. ER will run inside a terminal window and log messages will be shown there. Closing the terminal window will exit ER. If you want ER to always open a terminal window, modify the ER desktop shortcut created on installation and add/consoleto the end of theTargetline (outside the quote marks).
Task modules
- ER tasks are written in Python and based on the open-source SNAP framework
originally written by Christian Kothe and later extended by Intheon. ER comes with a set
of standard tasks already written which can be reconfigured by creating modified
.cfgfiles which reference those modules. You can also easily write your own tasks using the original SNAP framework. You can use the task modules shipped with ER, found in thesrc/modulesfolder of your installation, as a guide or template for writing your own tasks, or copy and adapt the modules to make variants of those tasks. All modules must be placed in thesrc/modulesfolder of your ER installation to be used by ER.
Signal quality checks
- You can optionally click "Signal Quality Check" prior to a session to showing various quality measures of the live signal. These can be useful to show that the cap is correctly positioned.
- If you have selected a CGX device as your source, you can do an impedance check (just as you can in the CGX Data Acquisition app), by clicking the 'CGX device check' button.
Session notes
- At the end of a task, a small "Session Notes" dialog will pop up where the operator can write a short note if necessary. Simply type the note and press enter to save/exit the dialog (no need to use the mouse to click the Save button). (Pressing the enter key without writing anything will not record any note.)
- The note is saved to the XDF file as an event marker prefixed with
session-notes-. - The notes dialog can also be activated at any time during a task by pressing the
F5key in the ER task window. It will be saved as an event marker to the XDF file with the timestamp of when the notes dialog was activated (not the time of when the note was note). This dialog can be activated multiple times in a session, as desired. - Since these session notes be saved in the XDF file, they should not contain any PII (otherwise the entire XDF file will be considered PII).
- Session notes are also saved to a text file on disk, in the same folder and with the same name as the XDF file, with a
notes.txtsuffix. (All notes from the same session are concatenated into the same file, which also contains the name of the task and the subject/session IDs for reference.) - In the event the ER session is aborted (by pressing
shift-ESCin the ER task window), the Session Notes dialog will also appear so that the operator can record the reason for the abort if desired. - The session notes dialog can be suppressed altogether by adding
show_session_notes_dialog: falseto the[Study]section of the.ersettings file used for the session. (The notes dialog will still be available with theF5key shortcut.)
Running Experiment Recorder in headless mode
- Experiment Recorder can also be run in "headless" mode. In this case, the main ER window will not pop up. You can run a single task by invoking a task config file directly, or you can run a series of tasks by invoking an .er file. This is useful for running an ER task from another script or application. To do, invoke the
experiment-recorder.cmdlauncher from the command line with the--nogui=trueargument. For example: experiment-recorder.cmd --settings=c:\studies\mystudy.er --nogui=trueto run the tasks inc:\studies\mystudy.er.experiment-recorder.cmd --module=c:\studies\tasks\mytask.cfg --nogui=trueto run a single task.- You can pass the subject_id, session_id and group_id (if needed) as arguments at the command line, i.e.,
experiment-recorder.cmd --module=c:\studies\tasks\mytask.cfg --nogui=true --group-id=group1 --subject-id=subject001 --session-id=1(note that for the command line arguments, the underscores have been replaced by dashes, sosubject-idnotsubject_id). - Note that starting with the 2023.1.0 release, the
--noguiargument will not take a value, it is simply--noguior-g.
Aborting a session
There are two ways you can abort a session in progress:
* Press shift-ESC in the task window (not the ER dialog).
* Call experiment-recorder.cmd --shutdown from the command line or another application.
In either case, the task is immediately terminated, the task window closed, the Neuropype recording pipeline is terminated (though the Neuropype service will continue to run in the background as usual and does not need to be restarted), and ER itself will shut down. To restart the session, launch ER in the same manner as before (i.e., via .er file, desktop shortcut, command line, etc.)
"Dry Run" mode
- You can run a task without recording to disk (i.e., for testing), by passing
--dry-run=trueas an argument if invoking ER from the command line, or by specifyingdry_run: truein your study.erfile.
Upload to Neuroscale
- If you have a Neuroscale account, ER can automatically upload your session files
to Neuroscale, if the
Auto-upload to Neuroscale after sessionoption is selected in theNeuroscalemenu (click on top-right icon to show the menu). If enabled, a dialog box will pop up at the end of the session showing the files to be uploaded, and uploading will begin automatically. - If you wish, you can cancel an upload by pressing the Cancel button in the Upload window when it appears.
- If there is no internet connection or Neuroscale cannot be reached an error message will appear and the upload will be cancelled. You can upload them later.
- You can manually upload your files to Neuroscale (i.e., if there was no internet connection at the time,
or you preferred to postpone the upload), by selecting
Neuroscale | Upload to Neuroscalefrom the ER menu (top-right corner). The upload dialog box will show all session files that have not yet been uploaded to Neuroscale, and automatically upload them. (If you cancel, the next time you runUpload to Neuroscaleyou will again get the list of files not yet uploaded, so this is not your only shot at uploading them.) - To upload files to Neuroscale, your Neuroscale storage credentials must be specified in
.erfile run (if no credentials are specified, an error message will appear). You can obtain your Neuroscale storage credentials by logging into your Neuroscale account atdashboard.neuroscale.ioselecting Storage, and clicking theCredentialsbutton.
Troubleshooting
- Some error messages you might get when running Experiment Recorder:
- Neuropype is not running. Run Neuropype by double-clicking the desktop shortcut. (If Neuropype is running and you get this message, exit Neuropype by right-clicking the N icon in the taskbar, and restart it.)
- LSL stream cannot be found. Make sure that your device is sending out an LSL stream, it is of type EEG, and it has the same name (case sensitive) as configured in your .er file (as indicated in the ER status bar). You can check the list of LSL streams by clicking the input source icon next to the status bar in the ER dialog. You have the option to abort the task, fix the issue, and start again, or "ignore" which will continue operation and perform the check again in 10 seconds at which time the error message will again appear if the LSL stream is still not found.
- A message appears while performing the task, that
no data is being recorded to disk. Possible reasons:- The LSL stream exists but the device is not sending data (i.e., battery is dead or connection is lost).
- You have a CGX device, your source is 'cgx', and it's sending data, but the device name does not match your selected device (your .er file says 'Quick-20r' but your device is 'Quick-20'; these two devices send data at different baud rates).
License
Experiment Recorder is released under the Neuropype license, which can be found in the Neuropype
folder of your installation (in the license.html file), and subject to the same terms as Neuropype itself.
Copyright (c) Syntrogi Inc. dba Intheon. All Rights Reserved.